RESEARCH PROJECTS
Under the umbrella of “Rare Disease Genomics in South Africa”, we use the latest in state-of-the-art “omics” and next-generation sequencing technologies to study African patients with genetic rare diseases. We are particularly fortunate to be able to offer African patients access to this technology, as they are understudied and under-represented in genomics worldwide. Our research group is housed within the new Biomedical Research Institute, which is a world-class facility -
watch the launch of the BMRI here
Our projects include:
The Undiagnosed Disease Programme in South Africa (UDPSA) – (see more info
here)
This is the first UDP in sub-Saharan Africa. We use whole exome sequencing and whole genome sequencing to find diagnoses for undiagnosed patients with rare diseases. Our current focus is on diagnosing paediatric patients as soon as possible, to be able to change medical treatment and management at the most impactful times. With time, we hope to include more adult patients. It is never too late to receive a diagnosis!
The results from the first year of the UDP have just been published in the American Journal of Medical Genetics Part A - of the first 100 patients, a diagnosis could be reached in 51 (a diagnostic yield of 51%). This is very encouraging, and shows that even in this understudied population of patients with rare diseases, exome sequencing has definitive utility. Read the paper here. For those still undiagnosed, we are exploring
different technologies and actively
searching for novel genes. Also, for this large project, we are building and optimizing the bioinformatic pipelines - we are particularly interested in students and postdocs who are keen to contribute to this effort - please email shahidamoosa@sun.ac.za
For the paper in the AJMG, Prof Shahida Moosa has been awarded the 2023 John M. Opitz Young Investigator award! Read more about the award here
Student projects within the UDP include exploring the genetic and genomic basis of several groups of disorders:
- Overgrowth
- Rare neurodevelopmental disorders
- Complex eye abnormalities
- Primary congenital glaucoma
- Adult neurological disorders
- Rare skeletal dysplasias and limb malformations
- Rare blood disorders
-----------------------------------------------------------------------------------------------------------------------------------------------------
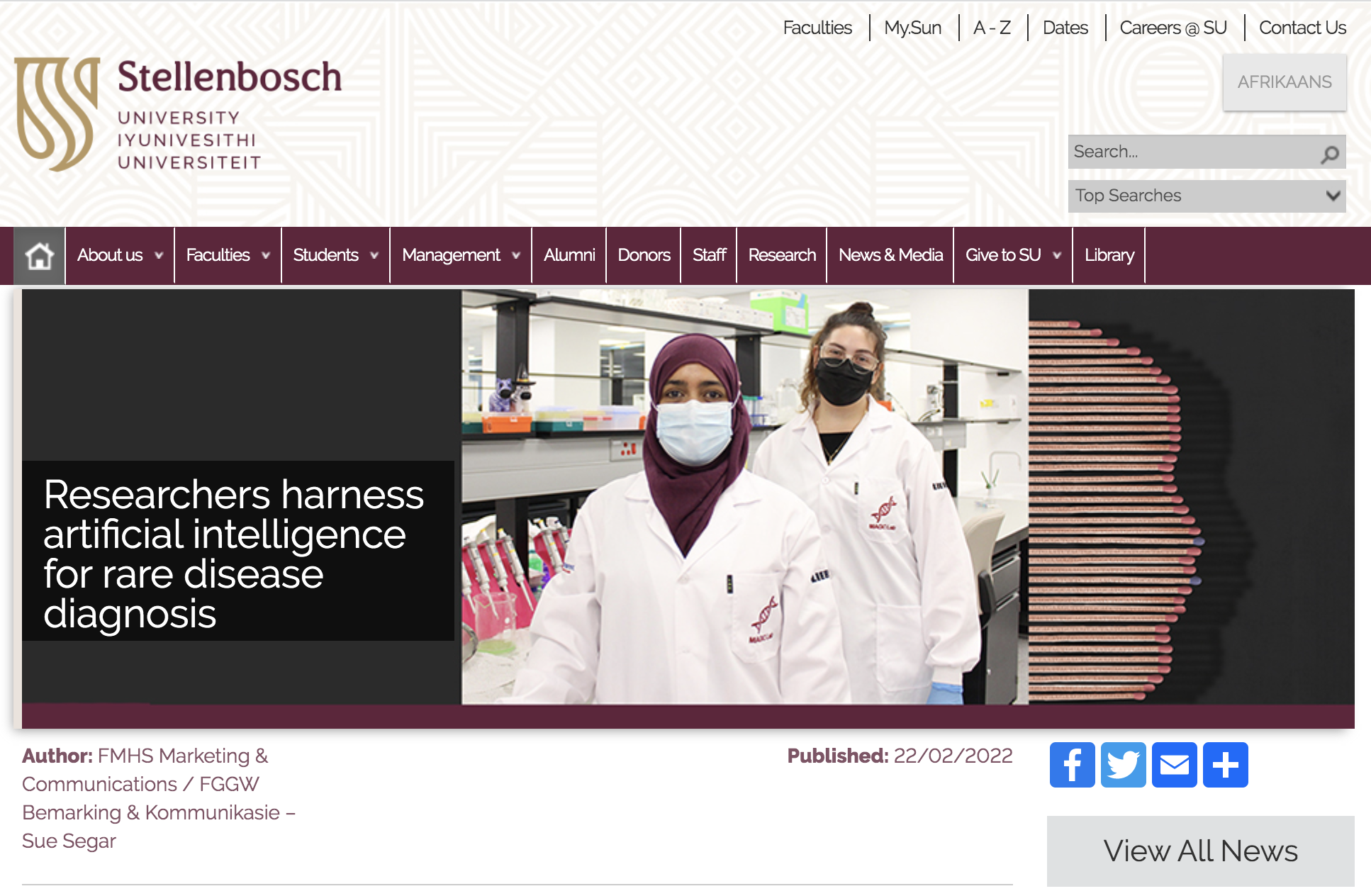
Next Generation Phenotyping using Artificial Intelligence - please see our
GestaltMatcher paper in Nature Genetics for work done in collaboration with the expert team of Prof Peter Krawitz at the University of Bonn in Germany. If you're interested in joining the
GMDB (GestaltMatcher Database community), please contact us. Read more about GMDB
here.
We have international collaborations looking at computer vs human vision in assessing images, to better understand how these processes differ, to enable improved training of both human trainees and computer-based learning.
-----------------------------------------------------------------------------------------------------------------------------------------------------
Genetics of Epilepsy across the lifespan in Africa - Epilepsy is the most common manifestation of neurological disease and the burden of epilepsy in Africa is significant. In children the prevalence of epilepsy is highest in the 1st years of life, and a genetic underlying cause is most often implicated. We conducted one of the first studies in Africa to determine the clinical utility of genetic testing in children with early onset epileptic encephalopathy. Using a next-generation-sequencing approach, we are able to provide almost half of our little patients with a diagnosis (paper published in Seizure). Early diagnosis and early treatment have a better impact on future cognitive and behavioural development. In addition, the genetic result plays a role in guiding physicians to decide on the type of anti-convulsant medication. Crucially, a definitive genetic diagnosis provides accurate information to parents on aetiology of the epilepsy and provides information for reproductive counselling for future pregnancies. There is paucity of data in Africa regarding epilepsy genetics and the genetic architecture underlying early onset epileptic encephalopathy in the African population.
To better understand the genetic contribution to epilepsy across the ages in Africa, a new pan-African project aims to recruit and test people with epilepsy. A genetic diagnosis can often guide treatment and other interventions, even in adults.
-----------------------------------------------------------------------------------------------------------------------------------------------------
Experiences of families with Rare Diseases - As very little is known about the lived experiences of families with Rare Diseases in South Africa, we have started to address this gap but exploring the experiences of caregivers, parents and siblings of people with rare diseases through qualitative research methods. This overarching theme comprises several projects aimed at looking at mental health, QoL and lived experiences of people with Rare Diseases in South Africa and their caregivers and family members, as a first step to be able to understand how to improve support. The work is done in collaboration with Prof
Chrisma Pretorius from the Department of Psychology at SUN.
-----------------------------------------------------------------------------------------------------------------------------------------------------
Health Economics related to Rare Diseases and genomic testing - Genomic testing is still considered "too expensive" for incorporation into routine clinical care. However, anecdotal evidence exists, for example from paediatric neurology, that earlier testing leads to considerable cost savings in the future. Therefore, we have several studies looking into the health economics of genetic and genomic testing in various clinics, starting with the impact of genetic testing in early onset epilepsy.
-----------------------------------------------------------------------------------------------------------------------------------------------------
The role of Genomics in the Paediatric Neurology Clinic: In addition to the study looking specifically at developmental and epileptic encephalopathy in our population of children, we have two further studies on the clinical utility of gene panels to diagnose paediatric-onset neurological conditions and on the utility of a "cerebral palsy" gene panel in children with CP. This is one area where we are able to directly influence management and even institute precision medicine. In real patients. In Africa.
-----------------------------------------------------------------------------------------------------------------------------------------------------Diagnosis of copy number variations using array-based comparative genome hybridisation in fetuses and neonates with congenital abnormalities in a South African population - Genome-wide assessment of copy number variations (CNVs) is recommended by numerous groups as a first-tier genetic test for the prenatal evaluation of fetuses with structural abnormalities detected on ultrasound and for neonates with unexplained multiple congenital abnormalities (MCA). Chromosomal microarray (CMA) is more commonly being utilized in prenatal and neonatal diagnosis due to its improved detection rate of clinically significant chromosomal abnormalities. This study aims to investigate the incidence of CNVs detected in fetuses with abnormalities on ultrasound and neonates with MCA in the South African population, in the hope to increase the diagnostic yield of genetic disorders using CMA.
-----------------------------------------------------------------------------------------------------------------------------------------------------
There are many more projects available, spanning clinical genomics, bioinformatics and molecular genomics as well as wet-lab molecular biology. Interested students and postdoctoral fellows as encouraged to contact Prof Moosa (shahidamoosa@sun.ac.za) to discuss possible projects.
-----------------------------------------------------------------------------------------------------------------------------------------------------
-----------------------------------------------------------------------------------------------------------------------------------------------------
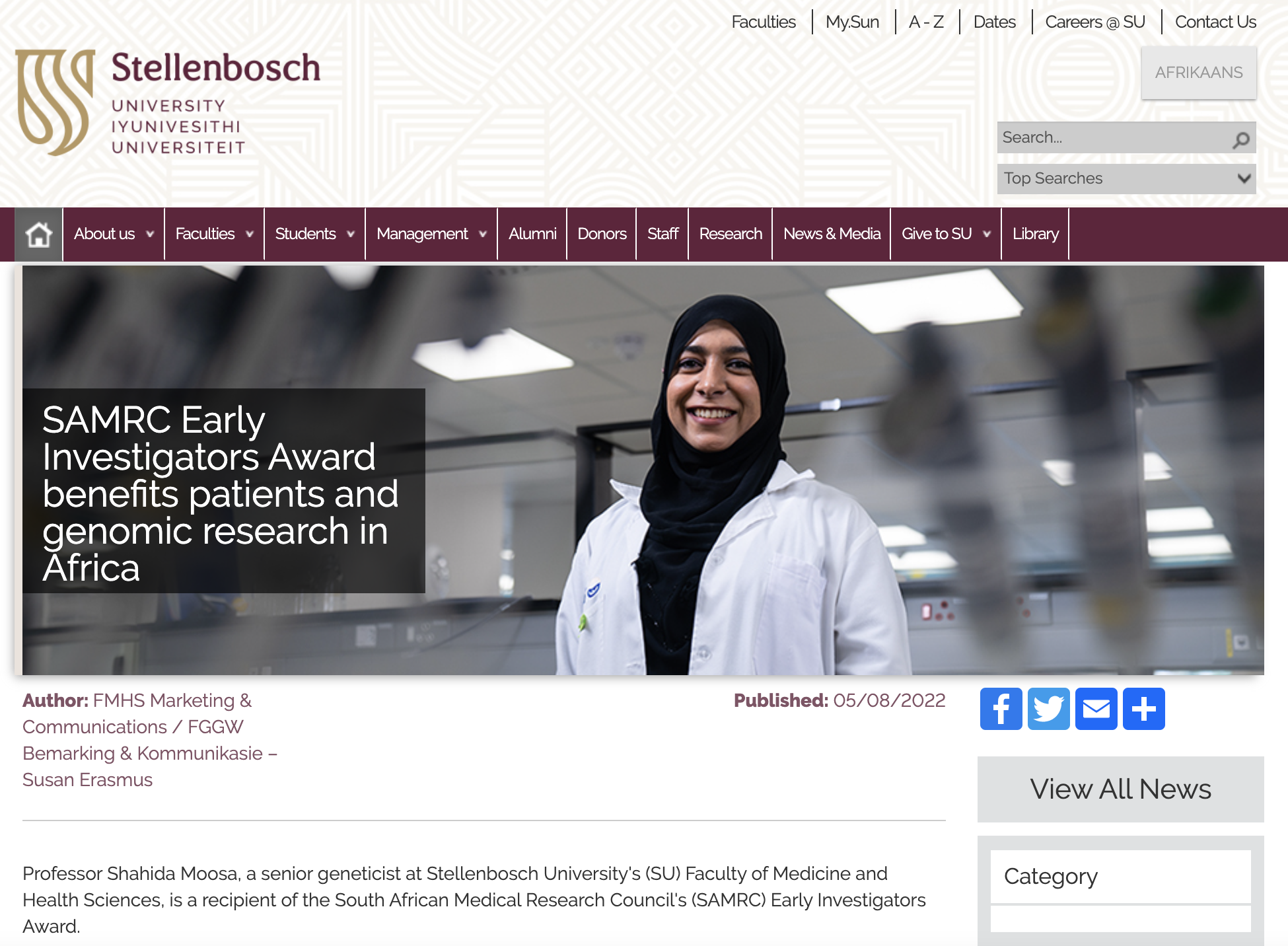
FUNDING
Along with international collaborations which have facilitated the research, Prof Shahida Moosa was recently awarded the SAMRC Early Investigator grant. This funding, totalling R2.5million, will go a long way to helping more families access genomics, and will primarliy support sequencing costs for the Undiagnosed Disease Programme.
Read more about the award
here